
Open GraMoFoNe plugin through the Plugins menu of Cytoscape.
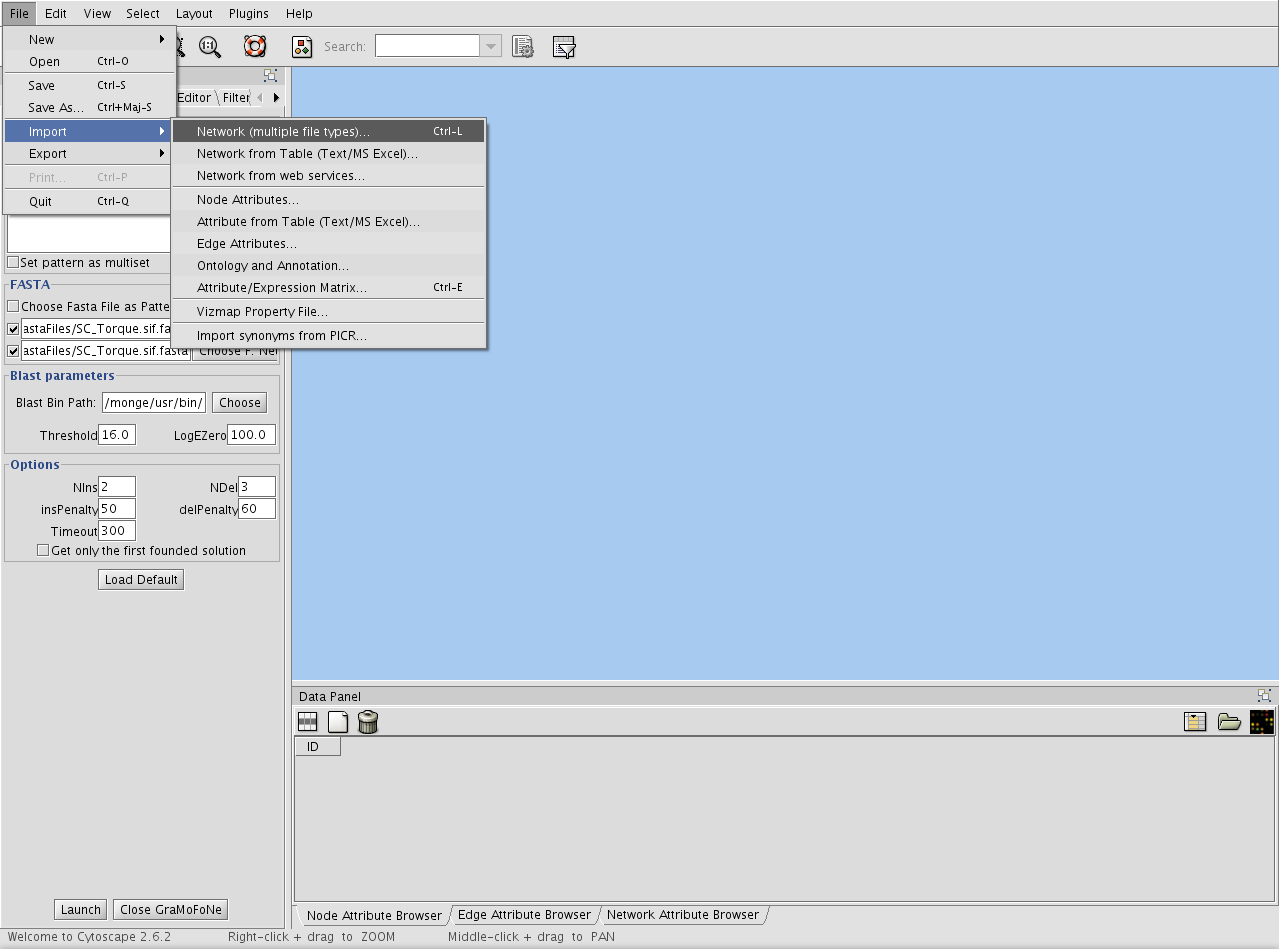
You must load a network to use GraMoFoNe. Open a network through the Import item in File menu.
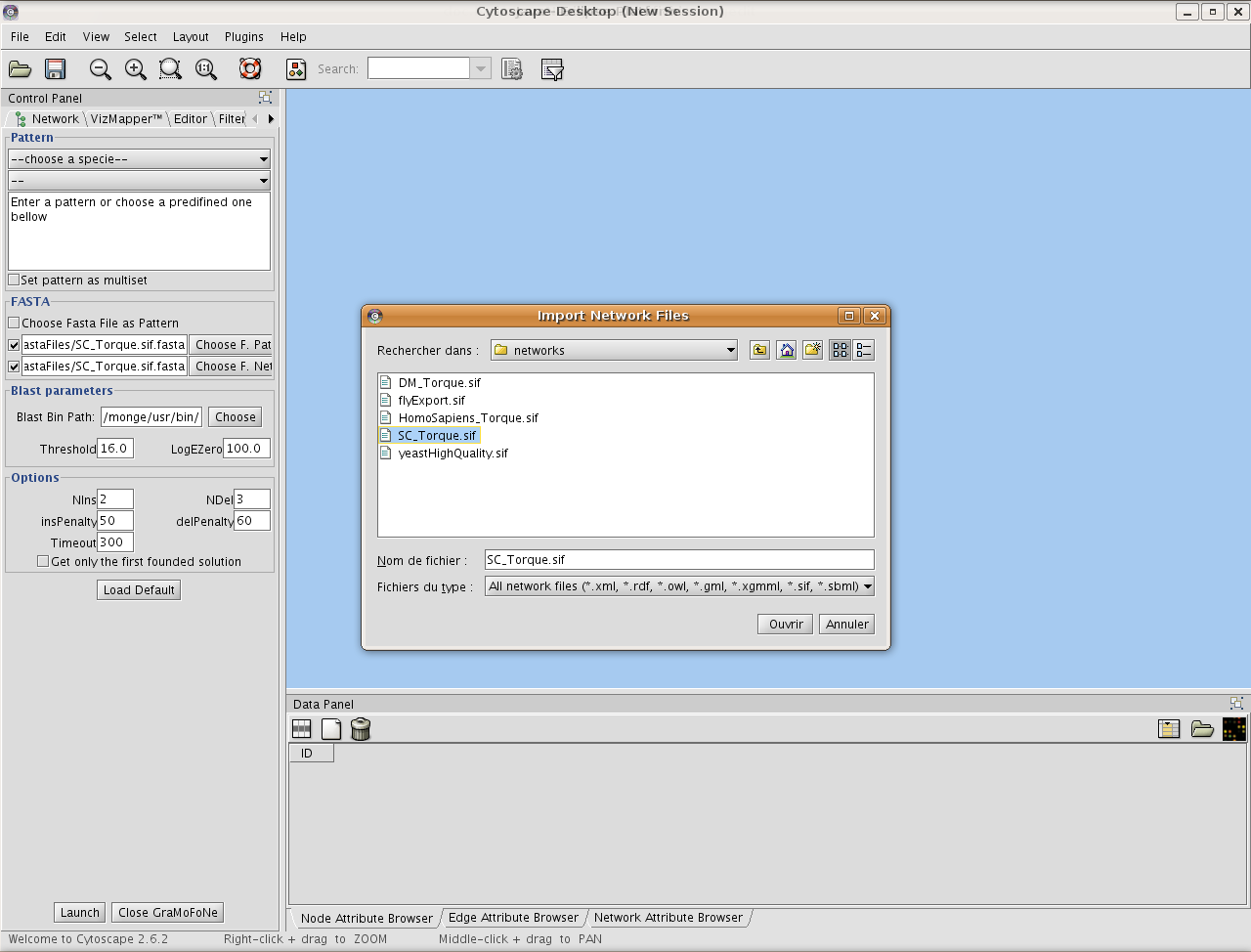
Choose a network format supported by Cytoscape.

Choose a motif to search into the network. You can type yourself the motif, choose one through the predefined ones or use your Fasta File as a motif. Here, we choose a predefined motif of the Yeast.
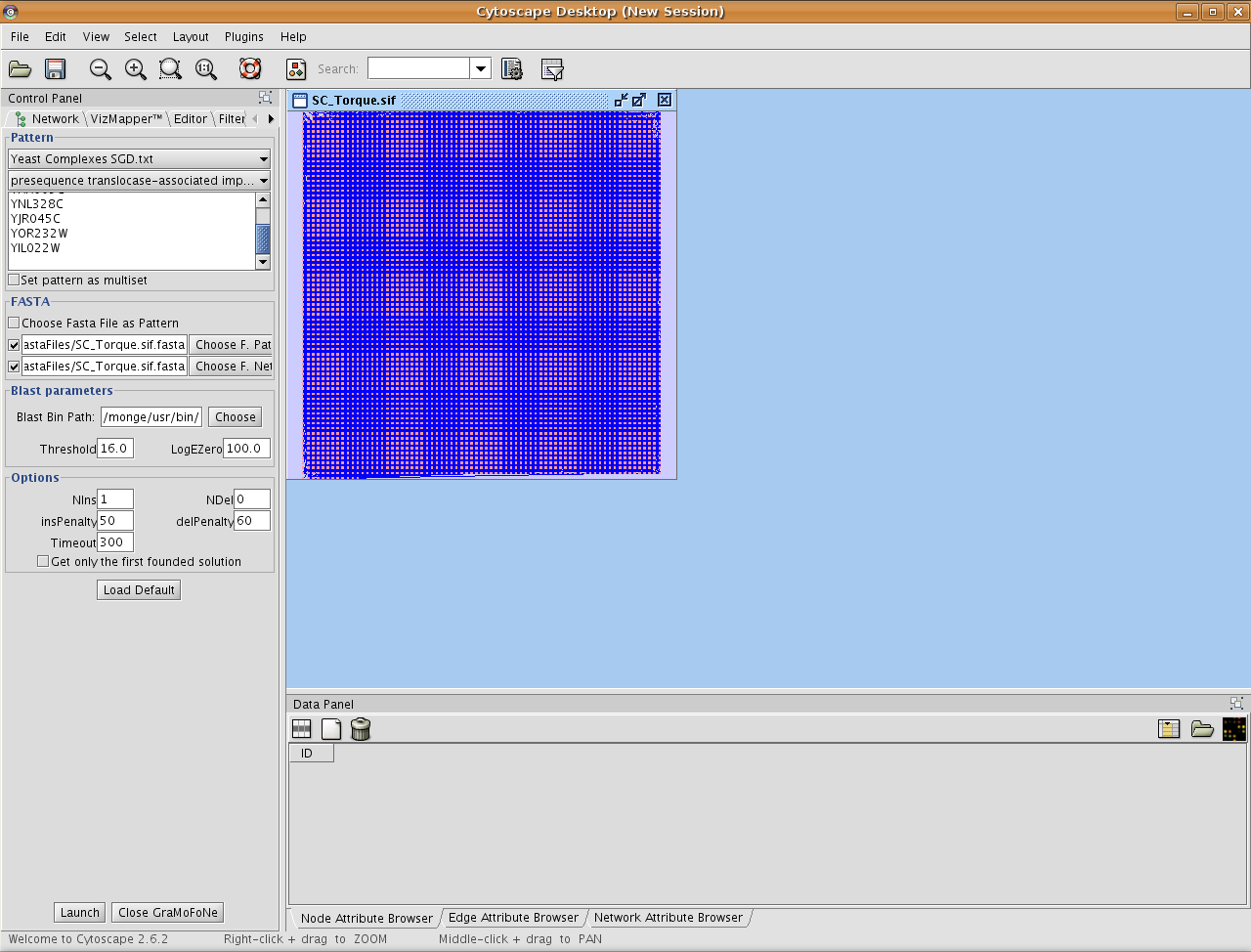
Choose your parameters for GraMoFoNe.
Pattern : You can set the pattern as a multiset (homologous proteins will have the same color).
Fasta : Choose the Fasta files for Motif (first one) and Network. If you don't check the box, the plugin will try to search the data on Uniparc database. You can check the box to choose the fasta file for pattern as a pattern.
Blast : You have to choose the path of you local Blast. You also can choose the threshold value to consider two proteins as homologous.
Options : You can choose the number of allowed insertions (proteins in the solution which was not in the motif) and allowed deletions (proteins in the motif and not in the solution). You can also choose the penalty for each one in the score function.
You can choose the timeout value. Until this time, the pseudo-boolean solver will stop.
Finally, you can choose to get the first solution founded by the pseudo boolean solver.

The plugin search occurrences of the motif in the pattern. You can cancel the task (stop after founding the next occurrence) without loss the previous occurrences founded.
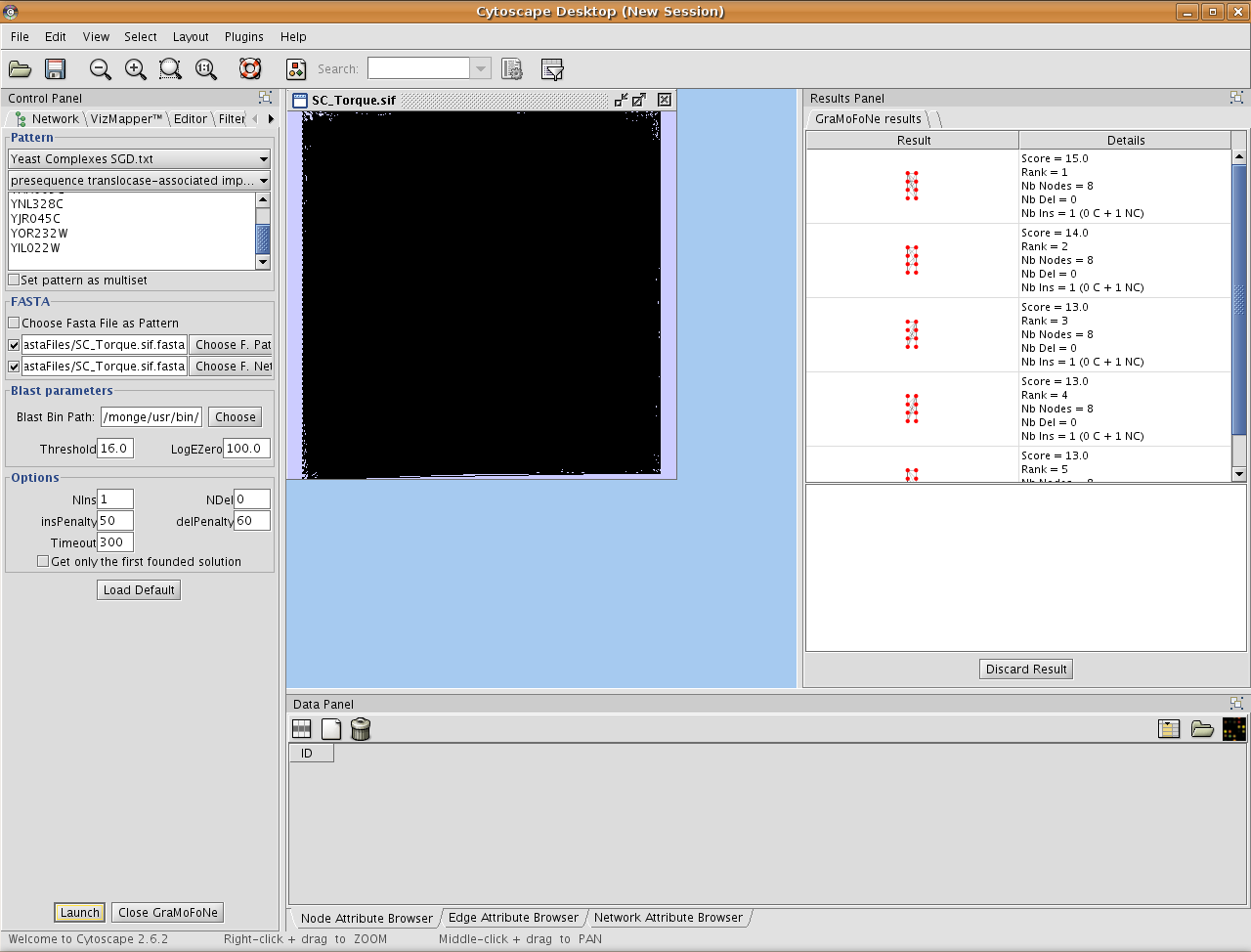
On the right side, there are all occurrences founded by our plugin.
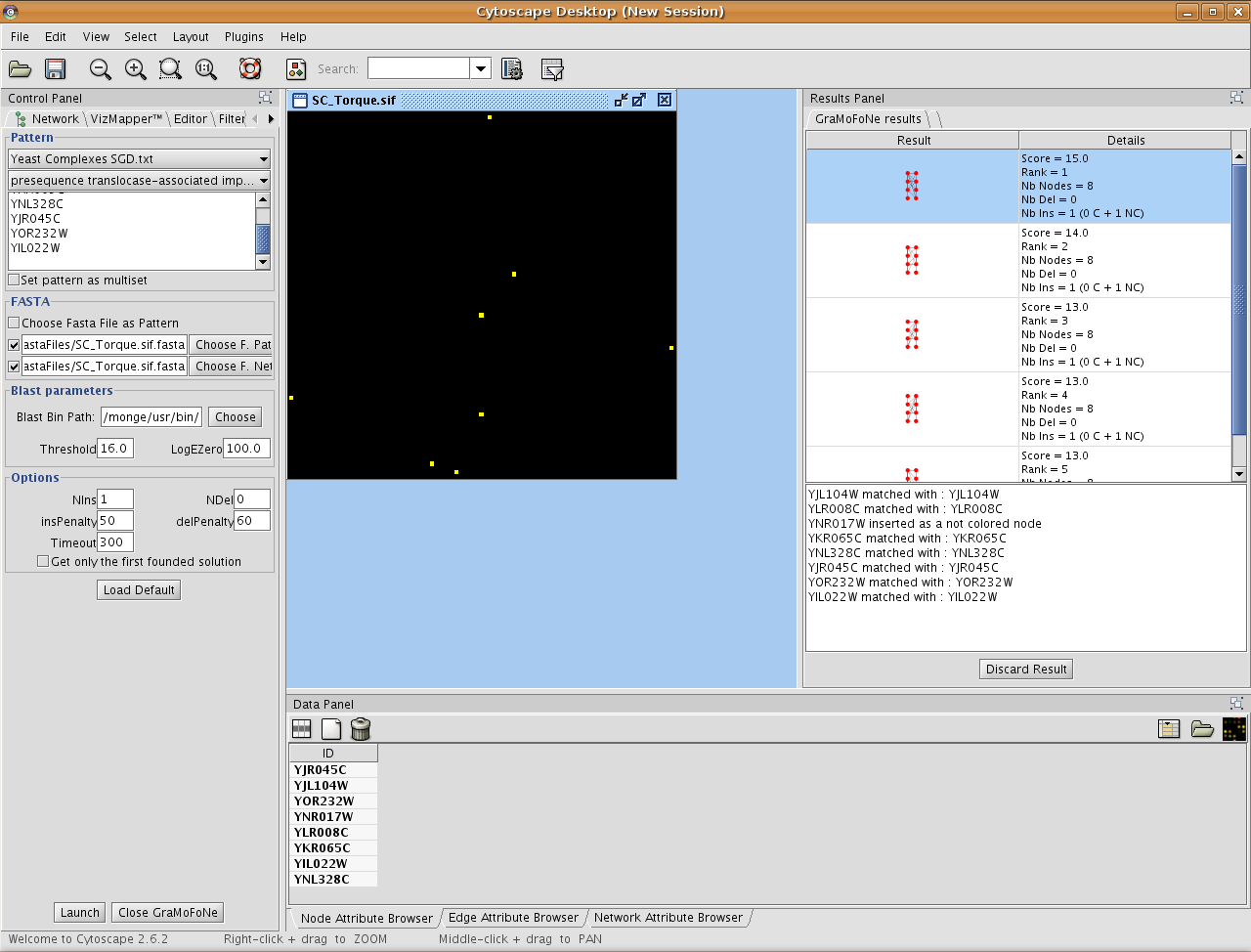
By clicking on one result, we highlight the occurrence in the network. Under the list of results, there is the correspondences between the motif and the proteins in the network.

By rolling the mouse, Cytoscape can zoom into the network.
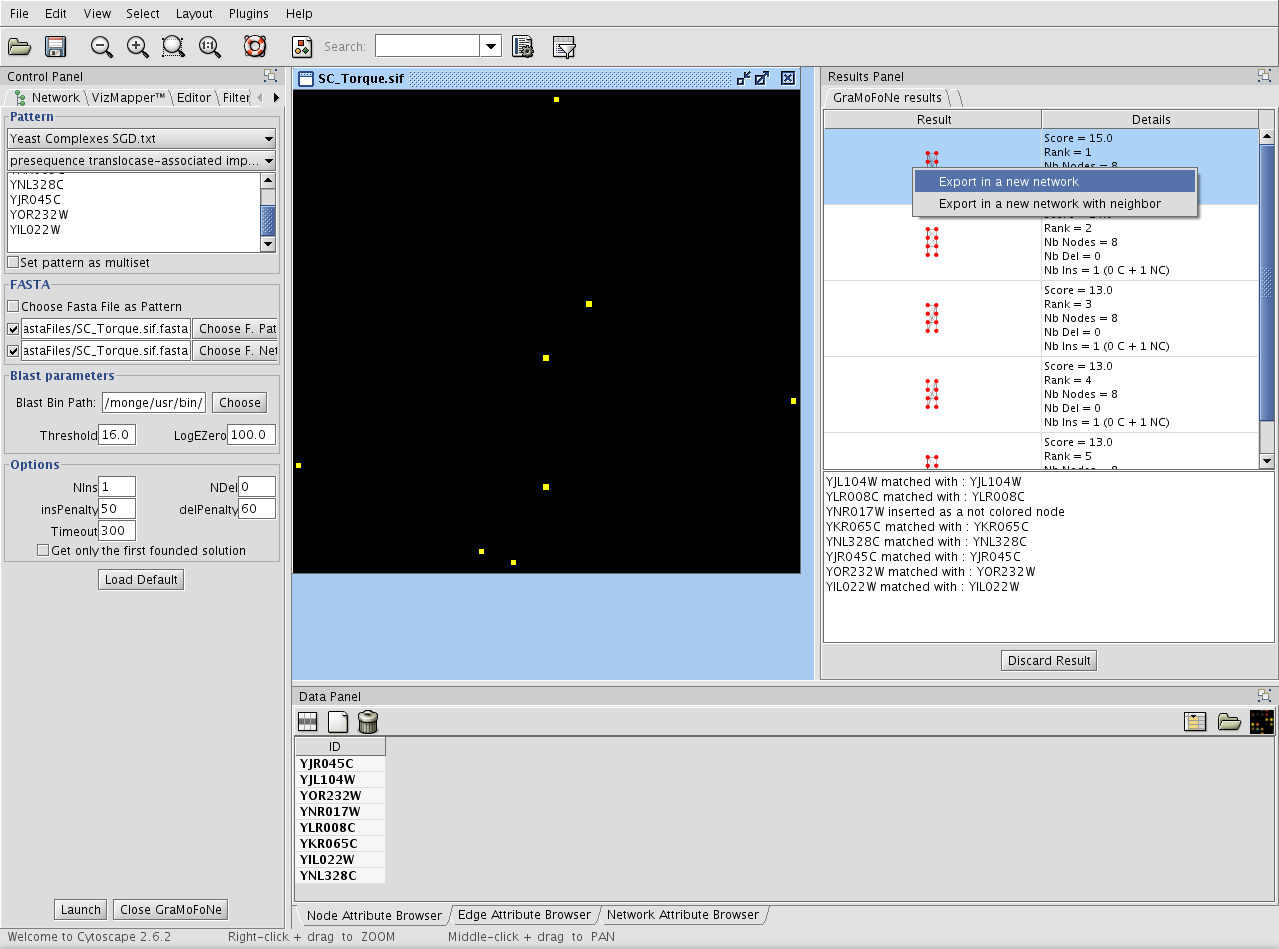
With a right click on the result, you can export the result as a new network. You can also export the result with all its neighbors.
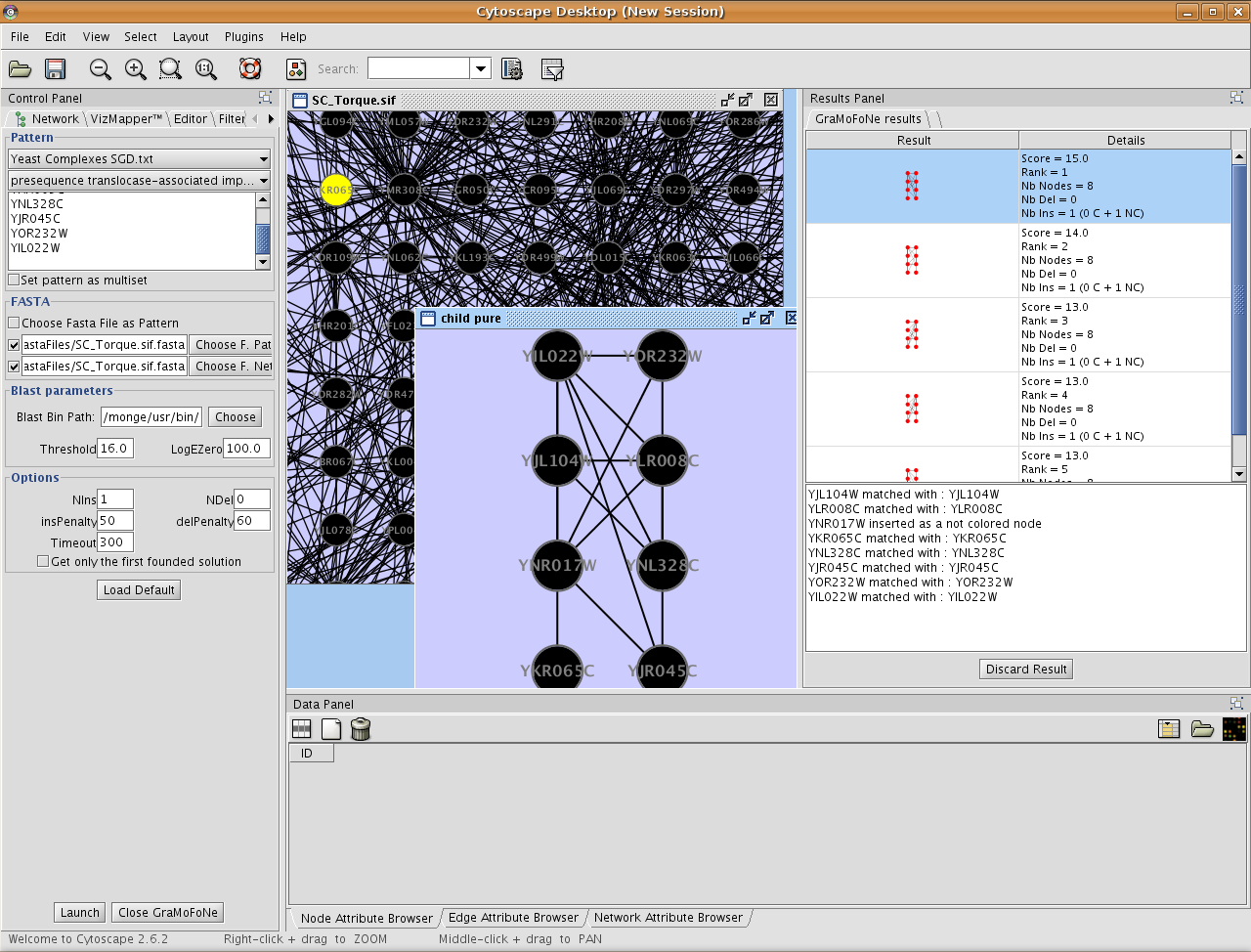
Exportation give a new network.
